Our work focuses broadly on asking questions about organismal function and evolution using genomic data. The huge amount of data currently being produced allows us to ask and answer questions on a genomic scale that have never been possible before. Our questions largely revolve around the relative roles of natural selection and genetic drift in shaping nucleotide, gene family, and gene expression variation both within and between species. Although most of the empirical work has been on systems such as humans, flies and mosquitoes (and now tomatoes!), members of the lab can work on topics and organisms that appeal to them. This page covers several major topics currently being studied.
The evolution of gene gain and loss
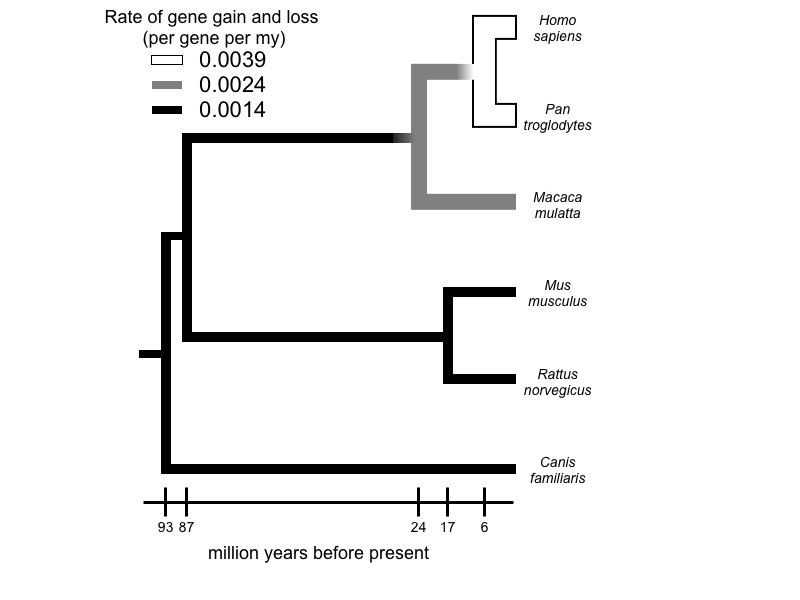
Comparison of whole genomes has revealed large and frequent changes in the size of gene families, the result of gene duplication and loss. Comparative genomic analyses allow us to identify large-scale patterns of change and to make inferences regarding the role of natural selection in gene gain and loss. To make these analyses possible, we have developed a stochastic birth-and-death model for gene family evolution, applied in the software package, CAFE. Application of this method to data from multiple whole genomes of many groups is revealing remarkable patterns of gene gain and loss. Other approaches to studying this question have involved the analysis of gene movement among chromosomes (especially sex chromosomes), the discovery of polymorphic copy-number variants under local selection, and even new methods for carrying out genome assembly to more accurately estimate gene number.
Population genomics
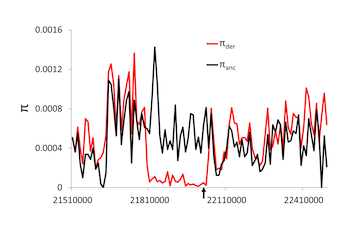
Selective, demographic, and random processes all determine the frequency of alleles in a population and differences between species. One of the major goals of population genetics has been to uncover which of these processes is acting in natural populations through a combination of directed empirical studies and theoretical models that provide expectations under a variety of conditions. While most of the work in the field has involved single loci or limited multiple locus studies and models, the availability of genomic-scale data will begin to require new genomic-scale approaches. We have been pursuing these questions in a wide variety of studies, largely focused on humans and flies (where the best data have always been). The work has presented new methods for distinguishing demography from selection, distinguishing different forms of positive selection, and more recently, using clinal variation to uncover local adaptation.
Speciation genomics
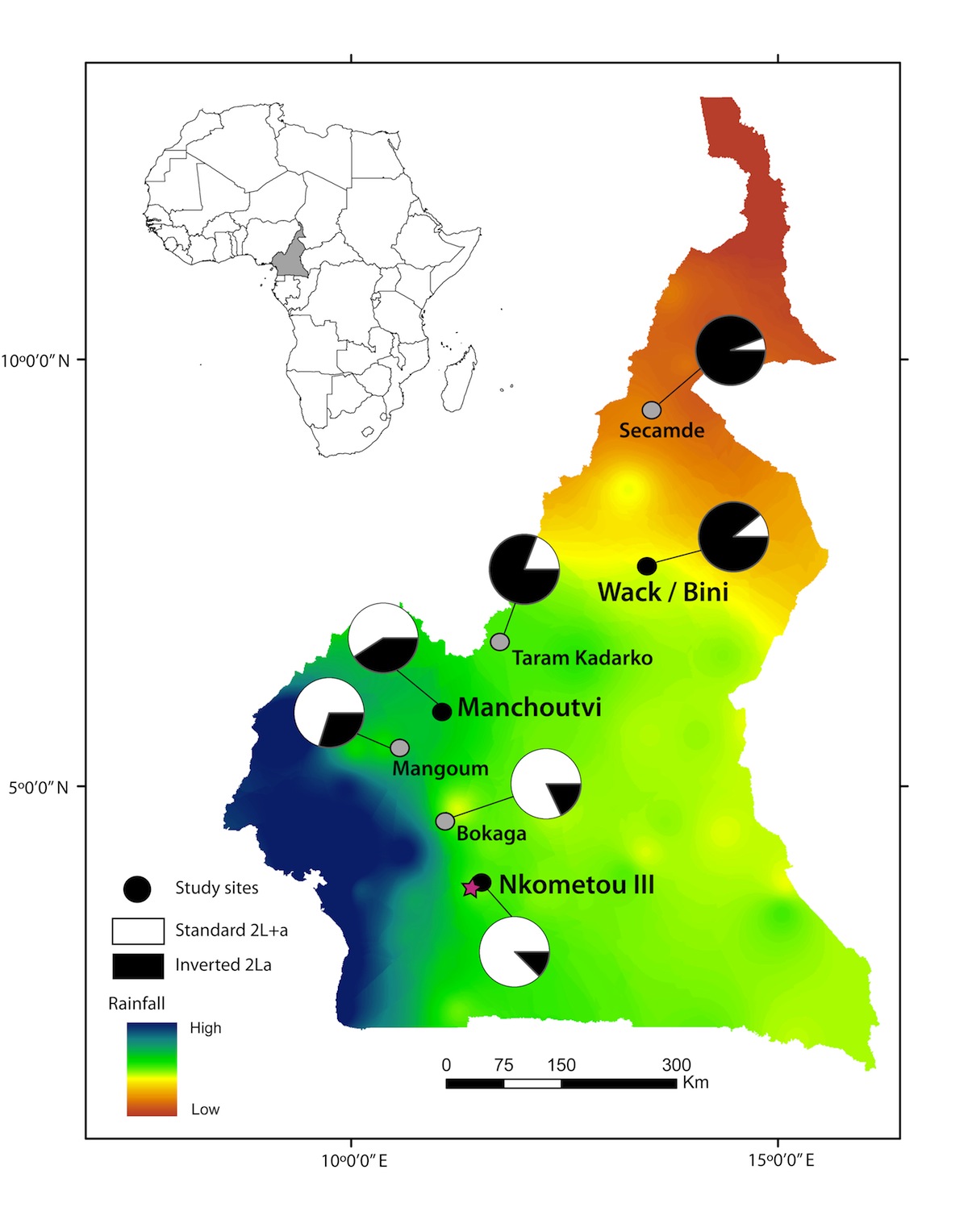
Population genomic data are being applied in new and creative ways to recently diverged lineages. One of the goals of the new field of speciation genomics is to understand how the patterns of divergence uncovered by such studies are related to mechanisms of reproductive isolation. Focusing on divergence within the Anopheles gambiae species complex, we have been interested in the roles of introgression and selection in shaping heterogeneous patterns of divergence. This work has built on much of our more general research into population genomics, but also now encompasses new approaches to detecting introgression and to distinguishing differences in introgression among loci from differences in selection among loci.
The evolution of transcriptional regulation
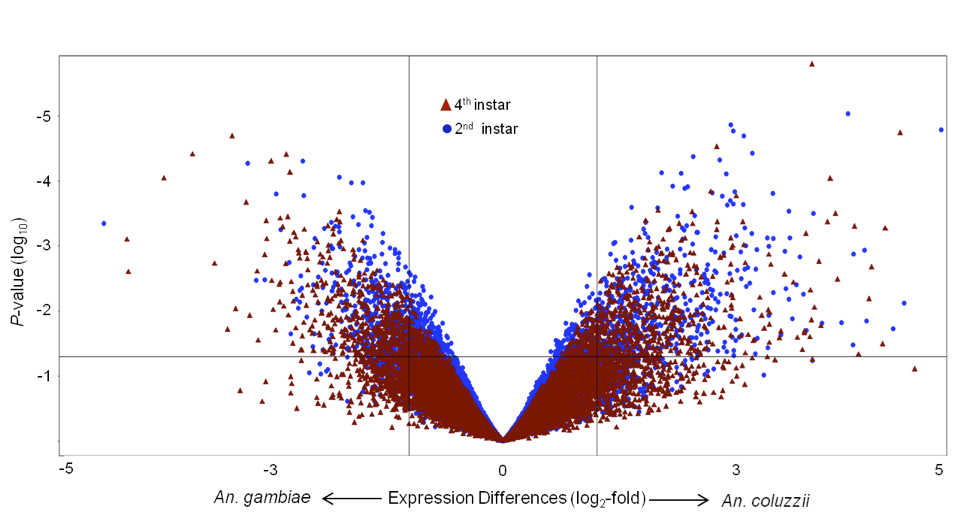
Changes in the timing, level, and location of gene expression have been implicated in many phenotypic differences between individuals and species. Using both DNA sequence and gene expression data, we can address the origin of variation in gene expression and the evolutionary forces that affect this variation. Our work has generally focused on the evolution of transcription factor binding sites and their implication for differences in expression, though with the rise of RNA-seq we are now more often than not starting with the observation of differences in expression. Much of the newer work uses wild tomatoes from the genus Solanum, as the ability to carry out genetic manipulations means that we can gain much more insight into the causes of transcriptional divergence.